In my research, I follow a philosophy of comprehensive understanding. Instead on focusing on a narrow taxonomical subset, I have extended my scope towards all lineages of venomous and poisonous animals and the totality of molecules that compose their chemical armory. I follow a step-wise research strategy, composed of five major stepstones that are outlined below:
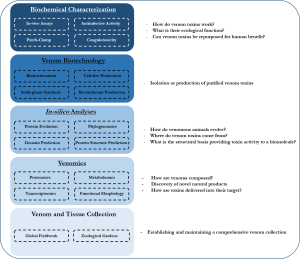
The stepwise research strategy employed by me and its major stepstones plus included methods within each (left) versus the questions asked and goals spearheaded towards (right).
On the most basal level of my work stands the mere venom collection. For the sake of establishing and maintaining a comprehensive toxin collection that emphasises material from rare and understudied taxa I am travelling the globe and conduct extensive fieldwork. I also cooperate closely with colleagues from zoological gardens across Germany.
The second level, and essentially the backbone of my work, is the venomics approach. By an integrative application of multi-omics (proteomics, transcriptomics and metabolomics) I decipher the chemical composition and gene expression of venom- and poison systems. The addition of functional morphology provides insights into the architecture of toxin delivery systems. In tandem, multi-omics and functional morphology generate a holistic understanding of the respective system from a chemical and biological perspective.
Next, I utilise bioinformatics to perform a variety of in-silico analyses. The combination of phylogenomics and protein evolution studies provides insights into the evolution of my focal taxa and their weaponised biomolecules. In parallel, the diversity of structural prediction, domain prediction plus other sequence analysis tools enables me to make structurally informed a priori assumptions upon the activity of a given toxin which can then be tested downstream.
Before testing of the gained assumptions becomes an option, the respective toxin needs to be obtained – which is the main focus of the fourth stepstone within my research strategy. While some toxins can be isolated via biofractionation, I commonly utilize venom biotechnology to produce them instead. This enables a consistent access to the produced toxin and allows for subsequent up scaling. Methods used to produce identified toxins are dependent upon the toxin type and range from solid-phase synthesis, cell-free approaches to recombinant expression in prokaryotes.
The fifth, and hence last, level of my research is the biochemical characterisation of the toxins. For this purpose a variety of bioassays for establishing the mode of action for a given protein are available. Assays included, are in vivo testing in different insect models (Galleria, Acyrthosiphon, Tribolium and Spodoptera), prokaryotes (e.g. MRSA), fungi, viruses and eukaryote cell lines. These are complemented with a series of in vitro assays, including patch clamp (in cooperation with colleagues), coagulotoxicity and enzyme activity tests. These assays provide insights into the activity and thus in the biological function of the studied toxin. Lastly, they allow to identify potential novel drug leads and enable the repurposing of natures most deadliest weapons for human benefit.